

Current version: 0.7.1
Note
By default, thefixespackage assumes time is a regularly spaced numeric variable (e.g., year = 1995, 1996, …).
If your time variable is irregular or non-numeric (e.g.,Datetype), settime_transform = TRUEto automatically convert it to a sequential index within each unit.
For unit-specific treatment timing, setstaggered = TRUE.
The fixes package is designed for convenient event study
analysis and plotting, particularly useful for visualizing parallel
trends and dynamic effects in two-way fixed effects (TWFE)
difference-in-differences (DID) research.
Key Functions:
run_es() — Takes a data frame, generates lead/lag
dummies, and fits the event study regression. Supports fixed effects,
covariates, clustering, staggered timing, weights, custom baseline, and
multiple confidence intervals.plot_es() — Plots event study results using
ggplot2 with flexible options: ribbon or error bars, choice
of CI level, and theme customization.plot_es_interactive() — Creates interactive event study
plots using plotly with hover tooltips displaying
coefficients, confidence intervals, standard errors, and p-values.Install from CRAN:
install.packages("fixes")Or with pak:
pak::pak("fixes")For the latest development version from GitHub:
pak::pak("yo5uke/fixes")First, load the library.
library(fixes)run_es() expects a panel data frame with at least:
Date)For staggered adoption
(staggered = TRUE), include a variable specifying
unit-specific treatment timing (e.g., “treatment_year”).
Widely used panel datasets include:
did::sim_dt(): simulated panel for DiD tutorialsfixest::base_stagg: a built-in dataset for staggered
adoptiondf1 <- fixest::base_did # Basic DiD
df2 <- fixest::base_stagg # Staggered treatment| y | x1 | id | period | post | treat |
|---|---|---|---|---|---|
| 2.8753063 | 0.5365377 | 1 | 1 | 0 | 1 |
| 1.8606527 | -3.0431894 | 1 | 2 | 0 | 1 |
| 0.0941652 | 5.5768439 | 1 | 3 | 0 | 1 |
| 3.7814749 | -2.8300587 | 1 | 4 | 0 | 1 |
| -2.5581996 | -5.0443544 | 1 | 5 | 0 | 1 |
| 1.7287324 | -0.6363849 | 1 | 6 | 1 | 1 |
| id | year | year_treated | time_to_treatment | treated | treatment_effect_true | x1 | y | |
|---|---|---|---|---|---|---|---|---|
| 2 | 90 | 1 | 2 | -1 | 1 | 0 | -1.0947021 | 0.0172297 |
| 3 | 89 | 1 | 3 | -2 | 1 | 0 | -3.7100676 | -4.5808453 |
| 4 | 88 | 1 | 4 | -3 | 1 | 0 | 2.5274402 | 2.7381717 |
| 5 | 87 | 1 | 5 | -4 | 1 | 0 | -0.7204263 | -0.6510307 |
| 6 | 86 | 1 | 6 | -5 | 1 | 0 | -3.6711678 | -5.3338166 |
| 7 | 85 | 1 | 7 | -6 | 1 | 0 | -0.3152137 | 0.4956263 |
run_es()The main event study function. All key arguments below:
| Argument | Description |
|---|---|
data |
Data frame to be used. |
outcome |
Outcome variable. Can be specified as a raw variable or a
transformation (e.g., log(y)). Provide it unquoted. |
treatment |
Dummy variable indicating the treated units. Provide it unquoted.
Accepts both 0/1 and TRUE/FALSE. |
time |
Time variable. Provide it unquoted. |
timing |
The time at which the treatment occurs. If
staggered = FALSE, this should be a scalar (e.g.,
2005). If staggered = TRUE, provide a variable
(column) indicating the treatment time for each unit. |
fe |
Fixed effects to control for unobserved heterogeneity. Must
be a one-sided formula (e.g., ~ id + year). |
lead_range |
Number of pre-treatment periods to include (e.g., 3 =
lead3, lead2, lead1). Default is
NULL, which automatically uses the maximum available lead
range. |
lag_range |
Number of post-treatment periods to include (e.g., 2 =
lag0 (the treatment period), lag1,
lag2). Default is NULL, which automatically
uses the maximum available lag range. |
covariates |
Additional covariates to include in the regression. Must be
a one-sided formula (e.g., ~ x1 + x2). |
cluster |
Specifies clustering for standard errors. Can be a character
vector (e.g., c("id", "year")) or a
formula (e.g., ~ id + year,
~ id^year). |
weights |
Optional weights to be used in the regression. Provide as a
one-sided formula (e.g., ~ weight). |
baseline |
Relative time value to be used as the reference category (default:
-1). For both classic and sunab
methods, this period is included in results with zero estimates.
Must be within the specified lead/lag range. |
interval |
Time interval between observations (e.g., 1 for yearly
data, 5 for 5-year intervals). |
time_transform |
Logical. If TRUE, converts the time
variable into a sequential index (1, 2, 3, …) within each unit. Useful
for irregular time (e.g., Date). Default is FALSE. |
unit |
Required if time_transform = TRUE. Specifies the panel
unit identifier (e.g., firm_id). |
staggered |
Logical. If TRUE, allows for unit-specific treatment
timing (staggered adoption). Default is FALSE. |
method |
Estimation method: "classic" (default) uses factor
expansion with fixest::i(), or "sunab" for
staggered-robust estimation using Sun & Abraham (2021)
decomposition. |
conf.level |
Numeric vector of confidence levels (e.g.,
c(0.90, 0.95, 0.99); default: 0.95). |
vcov |
Variance-covariance type (default: "HC1"). Accepts any
fixest::vcov() type (e.g., "HC3",
"CR2", "iid"). |
event_study <- run_es(
data = df1,
outcome = y,
treatment = treat,
time = period,
timing = 5, # Treatment occurs at period 5
fe = ~ id + period,
lead_range = 3,
lag_range = 3,
cluster = ~ id,
baseline = -1,
interval = 1,
conf.level = c(0.90, 0.95, 0.99)
)
# View summary
print(event_study)## Event Study Result (fixes)
## N: 1080 | Units: NA | Treated units: 1080 | Never-treated: NA
## FE: id + period
## VCOV: HC1 | Cluster: id
## Method: classic | lead_range: 3 lag_range: 3 baseline: -1fe must be a one-sided formula (e.g.,
~ firm_id + year).cluster can be a one-sided formula or a character
vector.# Basic plot with ribbon (default: 95% CI)
plot_es(event_study)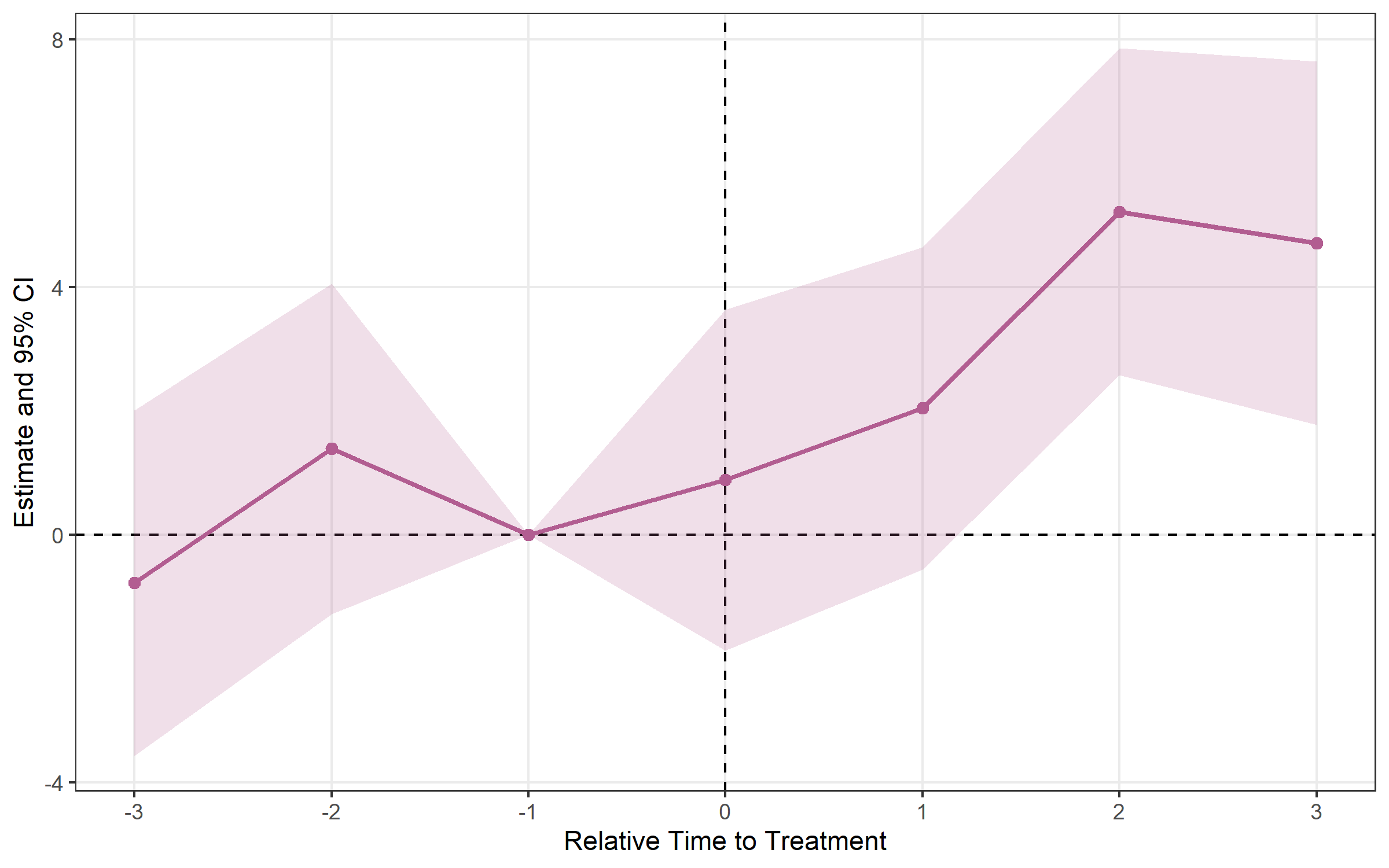
# Plot with error bars
plot_es(event_study, type = "errorbar", ci_level = 0.95)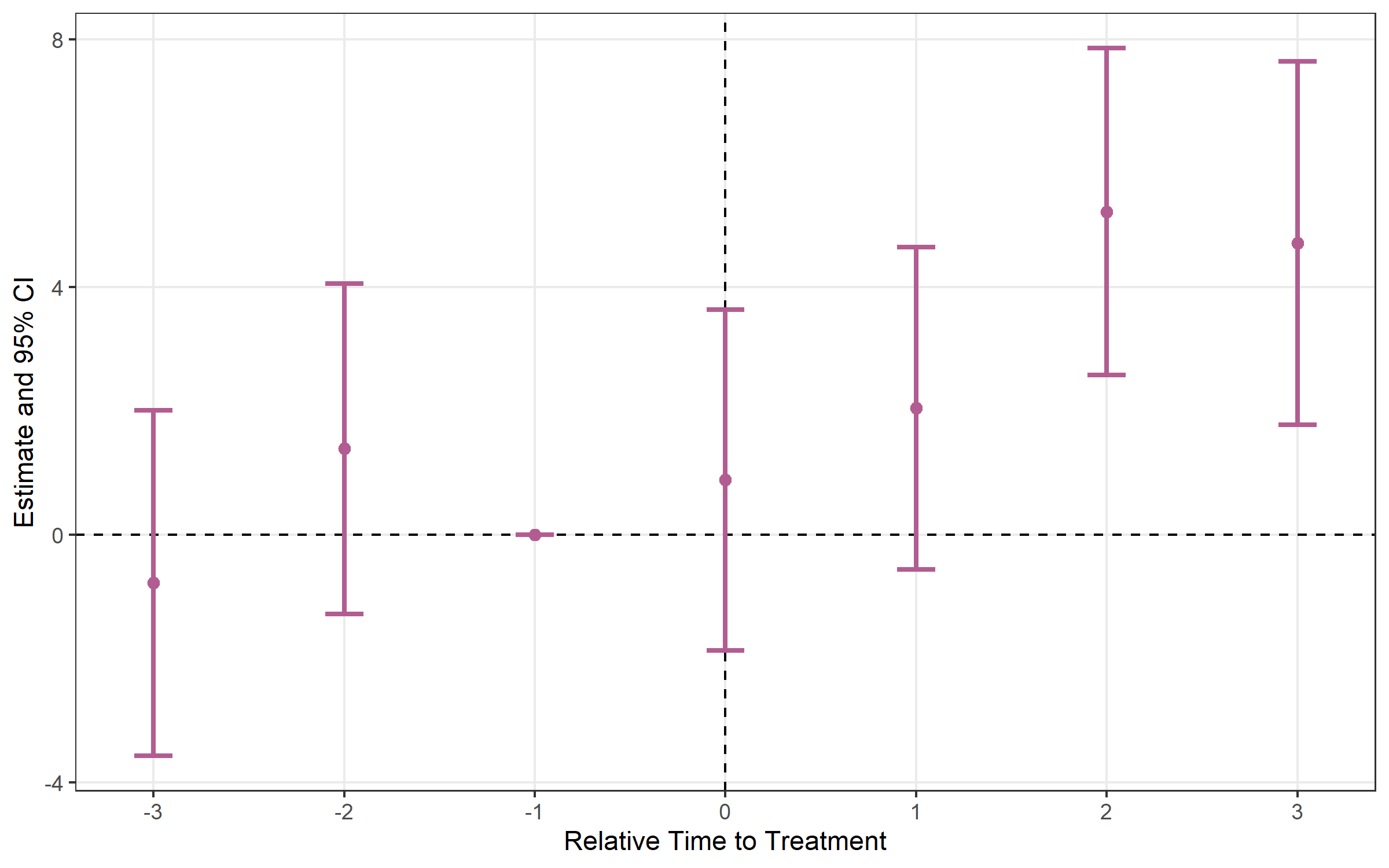
# Customized plot
plot_es(event_study, type = "ribbon", ci_level = 0.99, theme_style = "minimal") +
ggplot2::ggtitle("Event Study: 99% Confidence Interval")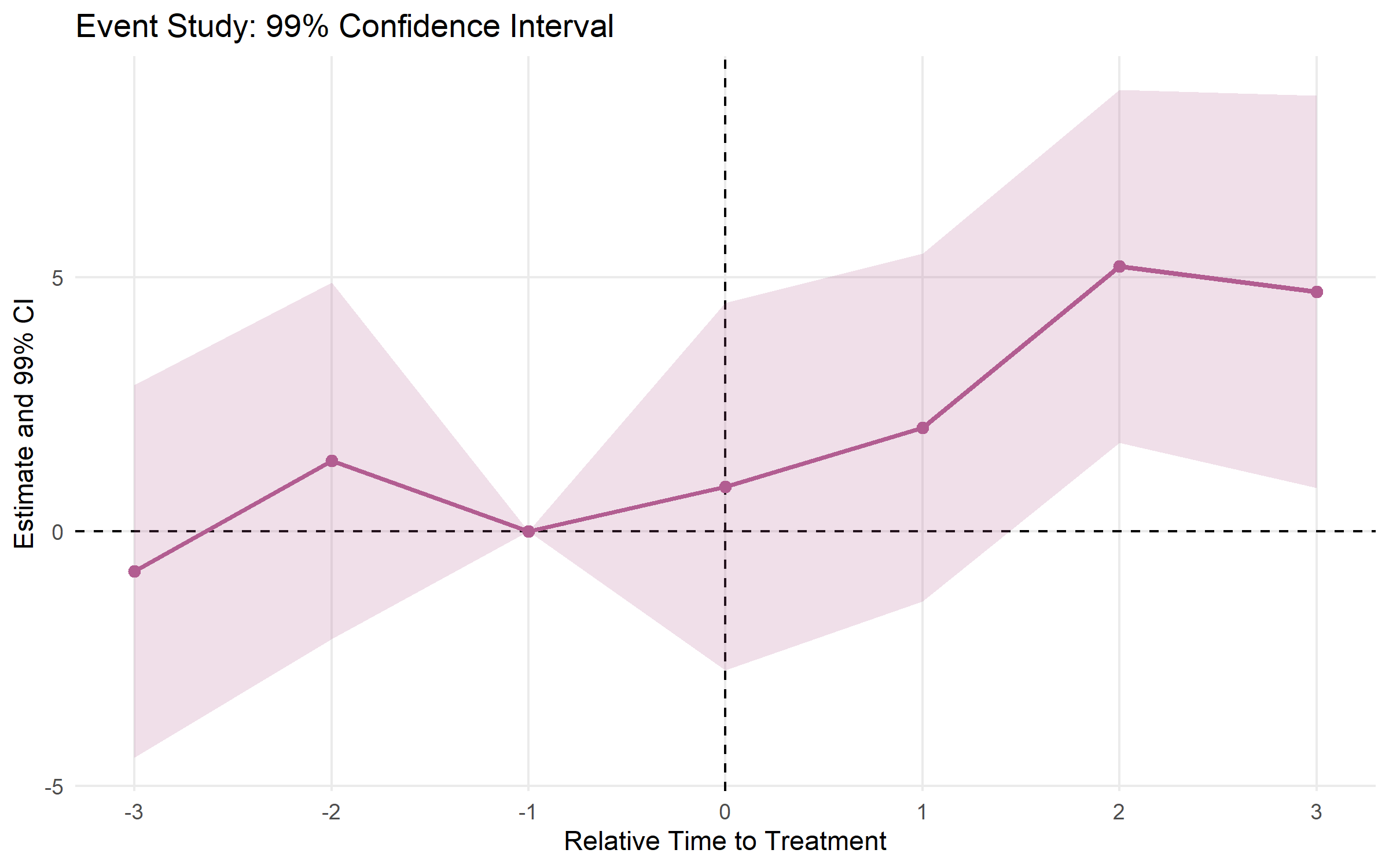
sunabFor staggered adoption designs with unit-varying treatment timing,
use method = "sunab" for the Sun & Abraham (2021)
estimator:
# Using fixest::base_stagg example data
event_study_sunab <- run_es(
data = df2,
outcome = y,
treatment = treated,
time = year,
timing = year_treated,
fe = ~ id + year,
staggered = TRUE,
method = "sunab",
lead_range = 3,
lag_range = 3,
cluster = ~ id
)
# View summary
print(event_study_sunab)## Event Study Result (fixes)
## N: 950 | Units: NA | Treated units: 950 | Never-treated: NA
## FE: id + year
## VCOV: HC1 | Cluster: id
## Method: SUNAB (staggered-safe)# Plot sunab results
plot_es(event_study_sunab)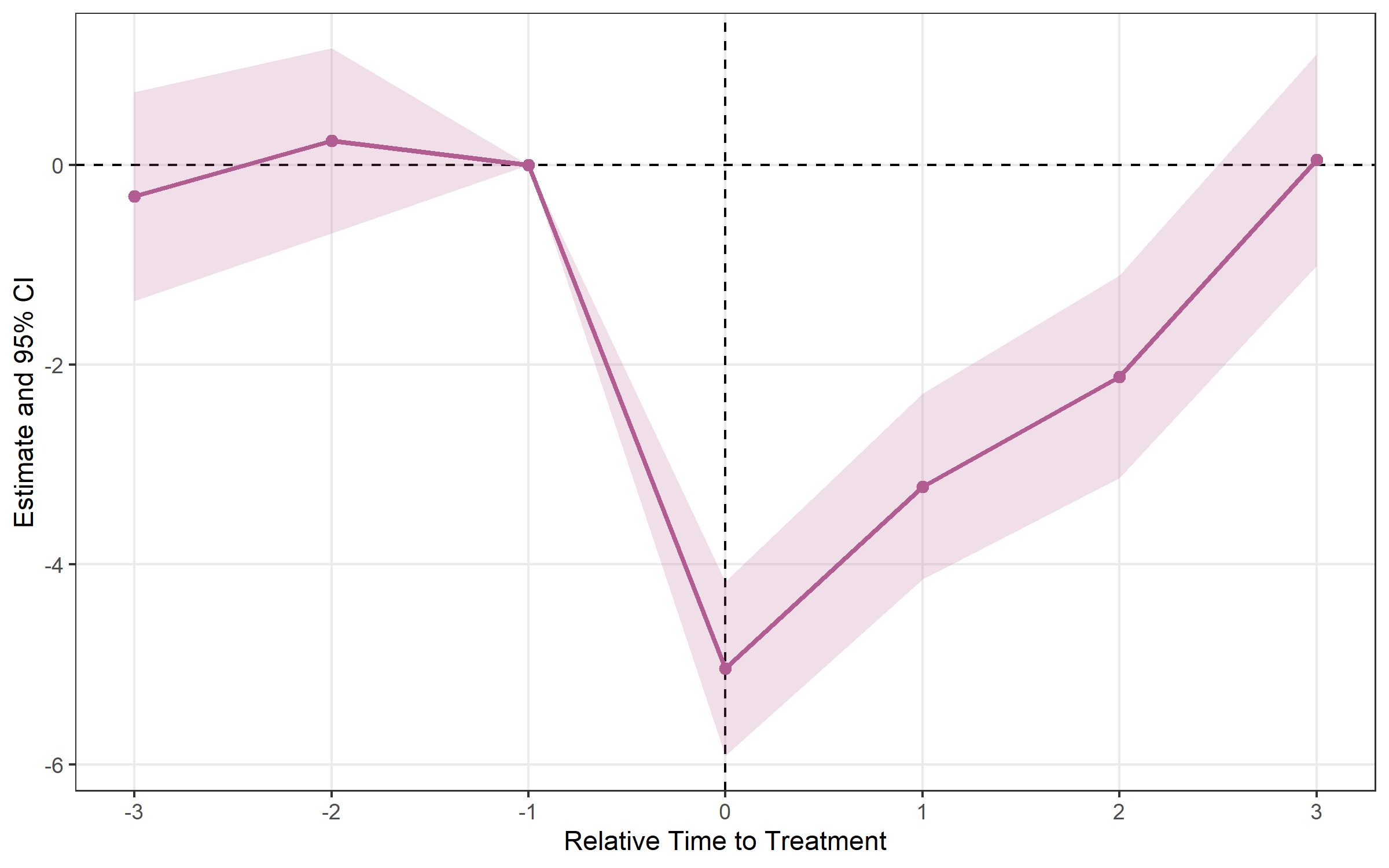
Note (v0.7.1+): The baseline parameter
now works for both classic and sunab methods,
and results are properly filtered to lead_range and
lag_range.
Create interactive event study plots with hover tooltips (requires
the plotly package):
# Interactive plot with hover information
plot_es_interactive(event_study, ci_level = 0.95)Hover over points to see relative time, point estimates, confidence intervals, standard errors, and p-values.
staggered = TRUE with
time_transform = TRUEtiming to accept original time values (e.g.,
Date), not just indexIf you find an issue or want to contribute, please use the GitHub Issues page.
Happy analyzing!🥂